Proteina-Complexa学习笔记
编辑1. Proteina-Complexa 介绍
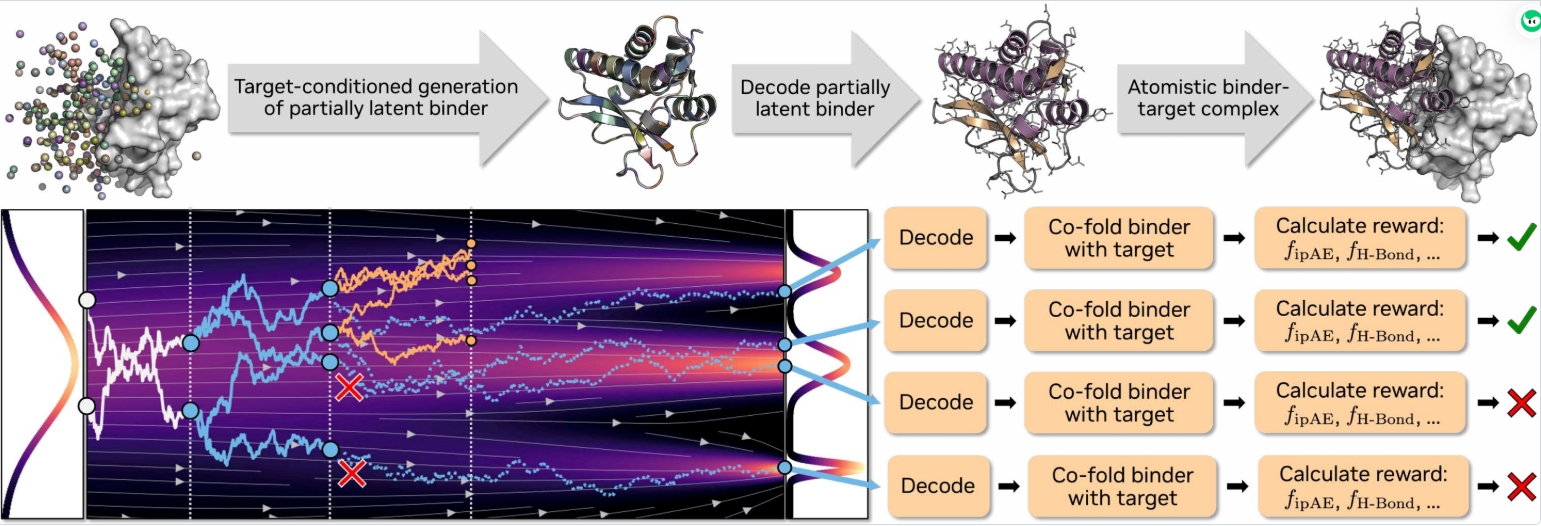
蛋白质相互作用建模是蛋白质设计中的核心内容。而机器学习技术的应用彻底改变了这一领域,使其在药物研发等诸多领域都发挥了重要作用。在这种背景下,基于结构的新型配体设计方法,可以被视为一种条件生成模型,或者通过结构预测工具来实现序列优化(“hallucination”)。Proteina-Complexa将两种方法进行结合,作者在现有的基于流的蛋白质结构生成技术基础上进行改进,利用通过计算方法预测得到的单体蛋白质结构之间的相互作用关系,构建了一个名为“Teddymer”的新大型数据集。该数据集包含了大量人工合成的配体-靶标对,可用于模型的预训练。结合高质量的实验用的多聚体结构,作者构建出性能出色的base model。随后利用这一生成的先验模型进行推理阶段的优化,从而将原本独立的生成技术和hallucination的优势结合。Proteina-Complexa在计算型结合体设计领域树立了新的标杆:与现有的生成式方法相比,它的计算机模拟成功率显著更高。此外,测试时间优化策略也使得其在相同的计算资源限制下,性能远远优于以往的各种方法。另外其还展示了界面氢键的优化方法、基于折叠结构的配体设计技术,以及将这些方法应用于小分子靶标和酶设计领域的应用案例。
2. Proteina-Complexa 安装
这里介绍docker安装
git clone https://github.com/NVIDIA-Digital-Bio/Proteina-Complexa
cd Proteina-Complexa
docker build -t proteina-complexa -f env/docker/Dockerfile .
运行,记得把参数换成自己的,默认权重会下载在protein-foundation-models目录的community_models和ckpts下面,因此我用git的文件夹替换了dockerfile内本身提供的protein-foundation-models,这样每次进行执行的时候能够获取权重
docker run --gpus all --rm -it -v /home/kangsgo/install/Protein_data:/workspace/data -v /home/kangsgo/install/Proteina-Complexa:/workspace/protein-foundation-models proteina-complexa
编辑.env_example文件,进行部分修改
# ==============================================================================
# Complexa Environment Configuration
#
# Setup:
# complexa init # Creates .env from this template
# # Edit .env with your values, then:
# complexa init <uv|docker> # Generates env.sh for your runtime
# source env.sh # Activates the environment
#
# WARNING: .env contains sensitive credentials.
# Do NOT commit .env to version control!
# ==============================================================================
# ==============================================================================
# USER CONFIGURATION — Edit these to match your setup
# ==============================================================================
# Credentials (SENSITIVE — fill in your values)
GITLAB_TOKEN=TOKEN_HERE
WANDB_API_KEY=YOUR_WANDB_KEY
WANDB_ENTITY=YOUR_WANDB_ENTITY
HF_TOKEN=
# Local paths (host-side) — set these to your machine's paths
LOCAL_CODE_PATH=/workspace/protein-foundation-models
LOCAL_DATA_PATH=/workspace/data/PFM_data
LOCAL_CACHE_DIR=${LOCAL_CODE_PATH}/.cache
LOCAL_CHECKPOINT_PATH=${LOCAL_CODE_PATH}/checkpoints
# Custom docker mounts (comma-separated "host_path:container_path" pairs, leave empty for none)
DOCKER_MOUNTS=
# Logging
LOGURU_LEVEL=INFO
# Cluster access
CLUSTER_USER=USER_NAME_HERE
# ==============================================================================
# DOCKER SETTINGS — Typically unchanged
# ==============================================================================
# Registry
REGISTRY=registry.example.com
REGISTRY_USER='$oauthuser'
# Docker image and container
DOCKER_IMAGE=registry.example.com/org/repo:tag
CONTAINER_NAME=proteina-dev
DOCKERFILE_PATH=env/docker/Dockerfile
# Docker-side paths (container-internal)
DOCKER_REPO_PATH=/workspace/protein-foundation-models
DOCKER_DATA_PATH=/workspace/data/PFM_data
DOCKER_PYTHONPATH=/workspace/protein-foundation-models/src
DOCKER_CHECKPOINT_PATH=/workspace/Proteina-Complexa/checkpoints
DOCKER_CACHE_DIR=/workspace/protein-foundation-models/.cache
DOCKER_HF_HOME=/workspace/protein-foundation-models/community_models/ckpts
DOCKER_HF_HUB_CACHE=${DOCKER_CACHE_DIR}/huggingface/hub
# ==============================================================================
# MODEL CHECKPOINTS — Derived from LOCAL_CODE_PATH, rarely need changes
# ==============================================================================
USE_V2_COMPLEXA_ARCH=False
# Community model checkpoints
COMMUNITY_MODELS_PATH=${LOCAL_CODE_PATH}/community_models
ESM_DIR=${COMMUNITY_MODELS_PATH}/ckpts/ESM2
AF2_DIR=${COMMUNITY_MODELS_PATH}/ckpts/AF2
RF3_DIR=${COMMUNITY_MODELS_PATH}/ckpts/RF3
RF3_CKPT_PATH=${RF3_DIR}/rf3_foundry_01_24_latest_remapped.ckpt
# ==============================================================================
# EXTERNAL TOOLS — Runtime-specific paths to tool executables
# ==============================================================================
# Python code reads the base names (FOLDSEEK_EXEC, SC_EXEC, etc.) via os.getenv().
# The base names default to UV paths. Change them to DOCKER_* if needed.
UV_VENV=${LOCAL_CODE_PATH}/.venv
# UV runtime tools (default for local development with .venv)
UV_FOLDSEEK_EXEC=${UV_VENV}/bin/foldseek
UV_RF3_EXEC_PATH=${UV_VENV}/bin/rf3
UV_SC_EXEC=${LOCAL_CODE_PATH}/env/docker/internal/sc
UV_MMSEQS_EXEC=${UV_VENV}/bin/mmseqs
UV_DSSP_EXEC=${LOCAL_CODE_PATH}/env/docker/internal/dssp
UV_TMOL_PATH=${UV_VENV}/lib/python3.12/site-packages/tmol
# Docker runtime tools (set in Dockerfile; also used for SLURM Pyxis)
DOCKER_FOLDSEEK_EXEC=/workspace/protein-foundation-models/bin/foldseek
DOCKER_RF3_EXEC_PATH=/workspace/.venv/bin/rf3
DOCKER_SC_EXEC=/workspace/protein-foundation-models/bin/sc
DOCKER_MMSEQS_EXEC=/workspace/protein-foundation-models/bin/mmseqs
DOCKER_DSSP_EXEC=/workspace/protein-foundation-models/bin/dssp
DOCKER_TMOL_PATH=/workspace/.venv/lib/python3.12/site-packages/tmol
# Active tool paths — Python reads these via os.getenv()
# Default to UV; change to ${DOCKER_*} to switch local runtime
FOLDSEEK_EXEC=${UV_FOLDSEEK_EXEC}
RF3_EXEC_PATH=${UV_RF3_EXEC_PATH}
SC_EXEC=${UV_SC_EXEC}
MMSEQS_EXEC=${UV_MMSEQS_EXEC}
DSSP_EXEC=${UV_DSSP_EXEC}
TMOL_PATH=${UV_TMOL_PATH}
DATA_PATH=${LOCAL_DATA_PATH}
# Active checkpoint path — YAML configs use ${oc.env:CKPT_PATH}
CKPT_PATH=${LOCAL_CHECKPOINT_PATH}
............
执行后执行如下命令下载权重
complexa download
随后可以初始化环境设置:
complexa init
complexa init docker
source env.sh
在执行目录下面新建一个bin文件夹
mkdir bin
cd bin
将https://github.com/cytokineking/FreeBindCraft/tree/master/functions
里面的dssp与sc下载并放入。
#也可以不做
wget https://mmseqs.com/foldseek/foldseek-linux-gpu.tar.gz
wget https://mmseqs.com/latest/mmseqs-linux-gpu.tar.gz
解压后放入bin目录下。至此安装完成。
验证是否安装成功:
# Validate the config resolves without errors
complexa validate design configs/search_binder_local_pipeline.yaml
3.快速开始
# 3. Design binders for PDL1
complexa design configs/search_binder_local_pipeline.yaml \
++run_name=pdl1_test \
++generation.task_name=02_PDL1
# 4. Check results
complexa status configs/search_binder_local_pipeline.yaml
其他管道类似如下:
# Ligand binder design
complexa design configs/search_ligand_binder_local_pipeline.yaml \
++run_name=ligand_test \
++generation.task_name=39_7V11_LIGAND
# AME motif + ligand binder scaffolding
complexa design configs/search_ame_local_pipeline.yaml \
++run_name=ame_test \
++generation.task_name=M0024_1nzy_v3
# Monomer motif scaffolding (indexed mode). Note motif targets not provided
complexa design configs/search_motif_local_pipeline.yaml \
++run_name=motif_test \
++generation.task_name=1YCR_AA
如果你和我一样,是24G显存,可以发现没法跑动,发现可以通过修改configs/pipeline/binder/binder_generate.yaml 中的dataloader batch_size改为8可以跑动。
TODO: 通过杨子辰老师的指导,主要是beam search和FK streeing费显存,可以进行修改。
- 0
- 0
-
分享